Intro
I’m sure most professional programmers by now have at least a passing familiarity with LLM-based coding assistants, whether through simple chat interfaces, or by trying not to get their prod data deleted by agentic approaches.
They’re not just a gimmick — they provide help with coding, apparent help with coding, and unambiguously deleterious "help" with coding. Key takeaway 💡: LLMs on occasion reduce cognitive load in 🧠 programming and OK, OK: I’ll stop with the oversaturated facsimile of stereotypical LLM writing patterns.
Regardless of what one might think about the current "AI" situation, coding agents, or even code-focused LLM chat assistants do help with the drudgery of programming on occasion.
Here, however, we’ll use both local and cloud-based LLMs to generate various outputs not traditionally thought of as "code": some to increase productivity or provide more options at work, and some to perhaps expand one’s overall toolset.
Before we proceed, one thing must absolutely be noted: for the topics covered, specialized models absolutely do already exist. The point here is not to show what can be done with modern models, but how already-familiar tools can be used in more ways, without any specialized knowledge, or additional deployment requirements.
Warmup: Making infographics
We’ll start with something more universally usable. On occasion, when communicating more complex messages to your coworkers/team, the text grows so rich in content, it becomes easy to lose the general view over the particulars. Which is where a graphical aid would come in handy – such as an infographic. But not everyone has the skills, patience, and/or time to create them, even when taking account the availability of various web tools meant for that purpose.
While it might seem counter-intuitive, generating graphics through a text-to-text coding-focused model is quite easy: just the matter of picking the right format; in our case, Scalable Vector Graphics (SVG).
SVGs are pretty much the most common files for vector graphics – primarily used for diagrams and illustrations – on the Web. The crucial bit about SVGs, however, is that internally, they’re just a completely human-readable XML file.
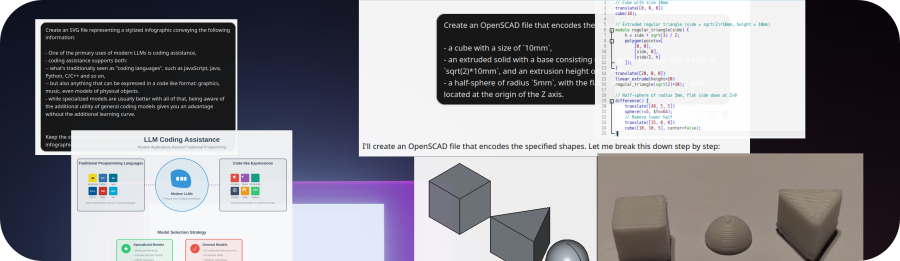
Here, let’s create a simple infographic for working with an LLM with the following prompt:
Create an SVG file representing a stylized infographic conveying the following information:
One of the primary uses of modern LLMs is coding assistance,
coding assistance supports both:
what’s traditionally seen as "coding languages", such as JavaScript, Java, Python, C/C++ and so on,
but also anything that can be expressed in a code like format: graphics, music, even models of physical objects.
while specialized models are usually better with all of that, being aware of the additional utility of general coding models gives you an advantage without the additional learning curve.
Keep the style clear, but in a typical infographic scheme. Enrich the infographic with small icons illustrating the most pertinent points.
Here’s one result, from Claude Sonnet 4.5 (through Copilot):

This was obtained from just a single prompt, no followups, and looks relatively good!
If you’re curious about the code itself, download the SVG file and open it in a code editor.
Local models don’t fare that well, comparatively, but still produce something approximating the desired result – here’s an example from GLM 4.5 Air after a couple of corrections:

While this has some formatting issues, it at least provides a valid approximation of what we’d want. And the great thing about generating these graphics in format such as SVG, as opposed to your typical genAI model outputting raster images, is that you can easily edit the file and correct any problems.
To wit, this is the same file after ~3 minutes of editing in Inkscape:

In short, generating SVGs with LLMs conveys a similar advantage as with programming language code: it provides a decent "jump off" point, which then can be changed, refined, and generally made to one’s liking; or just help with writer’s block, especially when not having significant experience with graphic design.
Main course: Prompts to objects
Intro
Now, for our main course: we’ll take the same chat assistants we use for coding, and make them help us create actual physical objects, just from a short concept spec.
Well, OK, for that to happen, we also require a 3D printer. For those unfamiliar with 3D printing: an extremely simplified procedure to go from design to physical object is as follows:
-
Design the object: typically in a CAD (Computer Aided Design) program, or via an equivalent coding framework (like OpenSCAD); alternatively, use a 3D editor like Blender.
-
Export the design to a format expressing surface geometry of the object(s), usually STLs nowadays.
-
Open the STL file in a slicer: a program that converts the objects surface geometry into an instruction sequence for the specific 3D printer setup (almost always split into layers, hence the name "slicer").
-
Send the instructions to the printer (here, we’re using an FFM printer), monitor the print, especially the first layer, wait for the print to finish, cool down (optional), remove the print, and clean it up as needed.
That first point reveals what we start with: we’re going to prompt the LLMs to create OpenSCAD specifications. Unlike most CAD programs, OpenSCAD is really just a language for specifying 3D models, plus a runtime with relevant functionality (rendering, exports, etc.). While some CAD suites do use human-readable formats, OpenSCAD is the most prominent one actually intended to be directly written and edited by humans.
The plan is as follows: we’ll start with using LLMs to attempt to describe some simple 3D solids, and, depending on the results, proceed to actually practical 3D prints.
Smoke tests
Let’s try a basic scenario to check if the LLMs do encode the ability to actually generate OpenSCAD specifications.
Here’s our starting prompt:
Create an OpenSCAD file that encodes the following:
a cube with a size of 10mm,
a regular triangle with a side of
sqrt(2),a half-sphere of radius 5mm.
We’ll start with cloud models, invoked via Copilot. Using ChatGPT-4.1, we obtain:
Here is an OpenSCAD file that creates the requested shapes. Each shape is positioned so they do not overlap.
// Cube with size 10mm translate([0, 0, 0]) cube(10); // Regular triangle with side sqrt(2) module regular_triangle(side) { h = side * sqrt(3) / 2; polygon(points=[ [0, 0], [side, 0], [side/2, h] ]); } translate([15, 0, 0]) regular_triangle(sqrt(2)); // Half-sphere of radius 5mm translate([30, 5, 0]) sphere(r=5, $fn=64); translate([30, 5, 0]) cube([10, 10, 5], center=true); difference() { translate([30, 5, 0]) sphere(r=5, $fn=64); translate([30, 5, -5]) cube([10, 10, 10], center=false); }The cube is at the origin. The triangle is placed at [15, 0, 0] to avoid overlap. The half-sphere is created using difference() to cut a sphere in half, placed at [30, 5, 0].
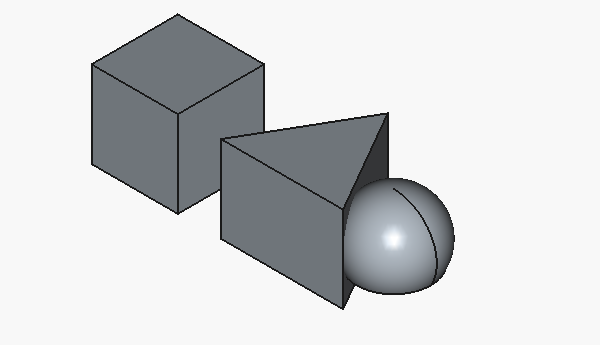
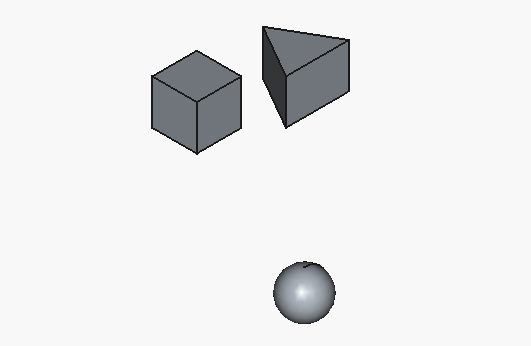
Aaand here’s how the result looks like:
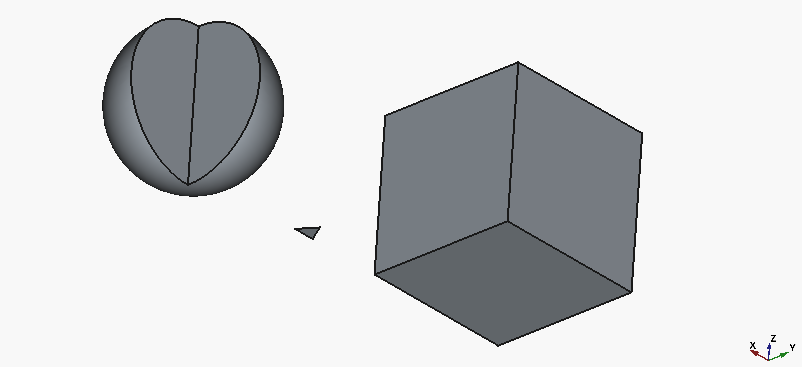
Well, we actually got working code, and that’s good… but only the cube looks half-way close to what we intended. To be fair, the triangle is on us, as we specified the parameters for a 3D object as if it was 2D.
Let’s give ChatGPT another go, with the following correction:
The triangle is considerably too small, and the half-sphere renders as a full sphere within a cube. Please fix.
| The "full sphere with a block" description actually refers to the presentation before a "cut" operation. |
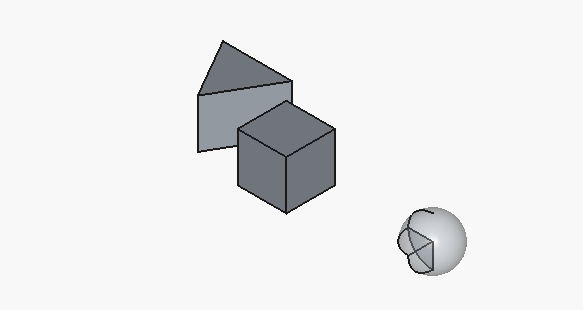
We obtain the following, supposedly corrected code:
Here’s a corrected OpenSCAD file: The triangle now has a more visible size (side length increased to 10mm). The half-sphere is properly rendered by subtracting the lower half of the sphere.
// Cube with size 10mm translate([0, 0, 0]) cube(10); // Regular triangle with side 10mm module regular_triangle(side) { h = side * sqrt(3) / 2; polygon(points=[ [0, 0], [side, 0], [side/2, h] ]); } translate([15, 0, 0]) regular_triangle(10); // Half-sphere of radius 5mm difference() { translate([30, 5, 5]) sphere(r=5, $fn=64); // Cut off the bottom half translate([25, 0, 5]) cube([10, 10, 5], center=false); }The triangle is now clearly visible. The half-sphere is created by subtracting a cube from the bottom half of the sphere.
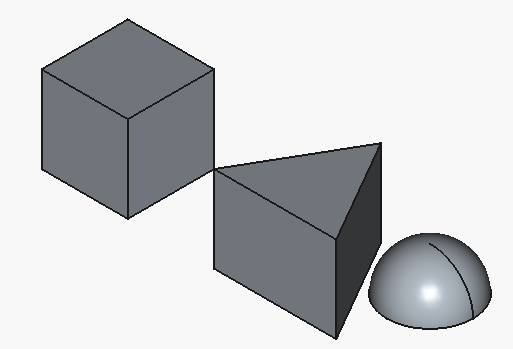
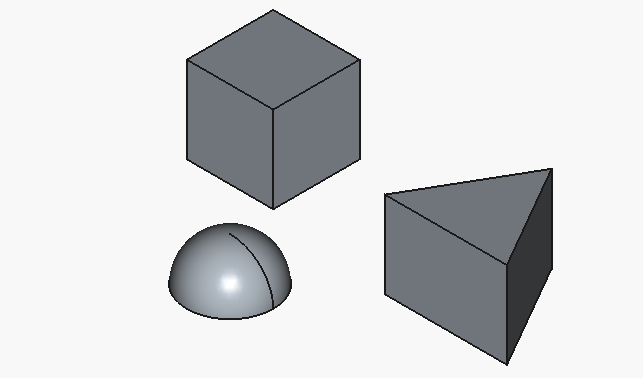
resulting in:
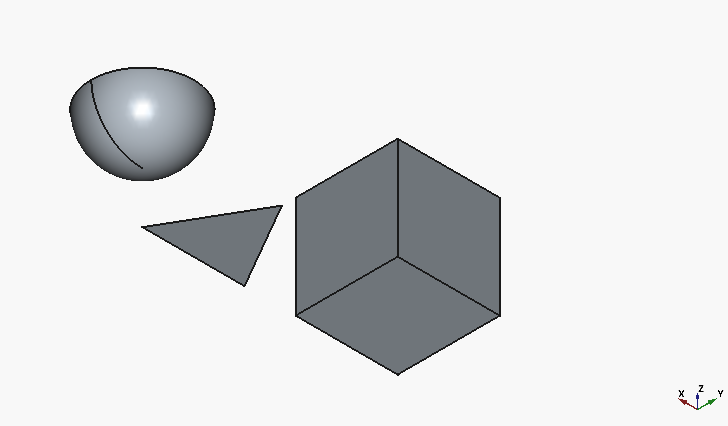
Much better, although, with all the hype towards the big-player LLMs, I would have expected for the half-sphere to be at least oriented with the flat face downwards.
Anyway, after asking once more:
Have the triangle extrude upwards with a face height of 10mm.
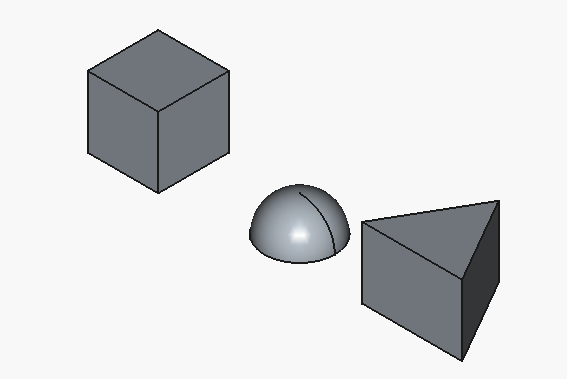
We can see the following:
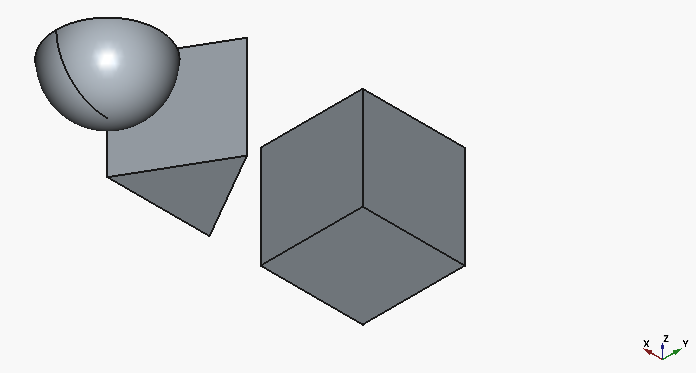
Let’s try to form a refined prompt that will get us to our target in one go:
Create an OpenSCAD file that encodes the following:
a cube with a size of
10mm,an extruded solid with a base consisting of a regular triangle with a side of
sqrt(2)*10mm, and an extrusion height of10mm,a half-sphere of radius
5mm, with the flat side facing downwards, and located at the origin of the Z axis.
The response is:
Here is an OpenSCAD file that creates the requested shapes, positioned to avoid overlap: Explanation: The cube is placed at the origin. The extruded triangle uses linear_extrude for a 3D solid, with the triangle base sized as specified. The half-sphere is created by subtracting the lower half of a sphere, with the flat face on the XY plane at Z=0.
// Cube with size 10mm translate([0, 0, 0]) cube(10); // Extruded regular triangle (side = sqrt(2)*10mm, height = 10mm) module regular_triangle(side) { h = side * sqrt(3) / 2; polygon(points=[ [0, 0], [side, 0], [side/2, h] ]); } translate([20, 0, 0]) linear_extrude(height=10) regular_triangle(sqrt(2)*10); // Half-sphere of radius 5mm, flat side down at Z=0 difference() { translate([40, 5, 5]) sphere(r=5, $fn=64); // Remove lower half translate([35, 0, 0]) cube([10, 10, 5], center=false); }This will generate the three shapes as described, each separated along the X axis.
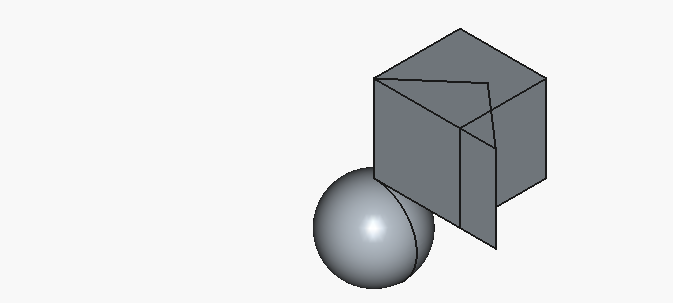
And the code renders to the following:
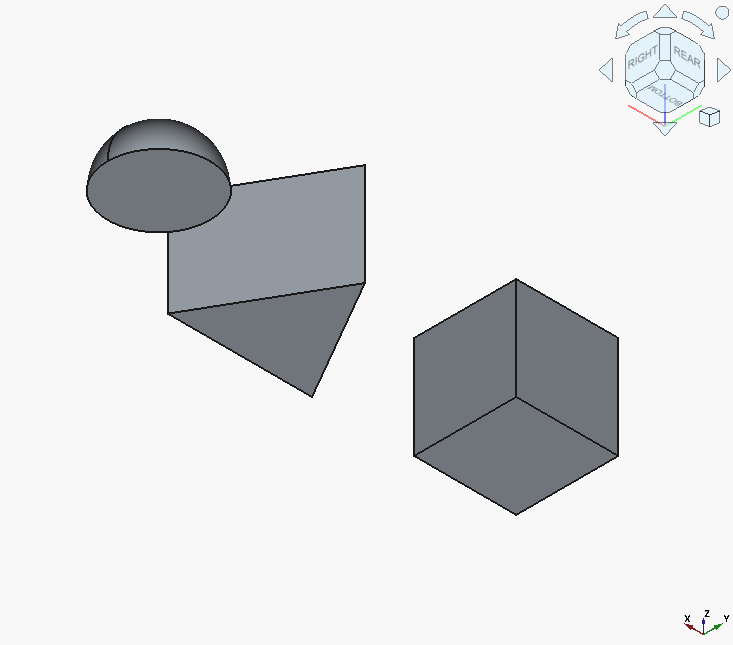
That looks about like something we wanted.
Baselines
Having established a baseline prompt, let’s run some tests comparing different LLMs. The selection of models stems from both what is available through Copilot, and what is currently thought of as "good" local LLMs tailored for code generation.
The latter was produced through an extremely rigorous and formal approach of an ensemble method composed of the result of a deep research query based on the currently expressed trends in the prosumer space, validated by a cross-reference from the immediately available wetware compute&storage system instance.
In other words: I ran the query on last month’s posts in r/LocalLLaMA and checked it wasn’t BS ,compared to my own impressions.
Local model selection
The models we’re going to use here are all unsloth’s quantizations/optimizations, specifically (name: variant):
-
Devstral-Small-2507: UD-Q8_K_XL
-
GLM-4.5-Air: UD-IQ2_XXS
-
Qwen3-Coder-30B-A3B-Instruct: UD-Q8_K_XL
-
gpt-oss-20b: F16
As far as configuration goes:
-
everything runs through llama.cpp (with llama-swap for convenience),
-
for every model, the context window size is 16k (mostly for a "thinking buffer"),
-
--jinja(unsloth’s suggested pre-prompts) is enabled wherever recommended in the model card.
A final clarification: the specific choice of quantization originates from the goal of all models fitting in roughly the same amount of VRAM (under 48 GB). This is theoretically not strictly "fair"[2], but, firstly, it matches actual usage scenarios better, and modern larger models often exhibit surprisingly good performance even with more aggressive quantizations anyway.
Setup
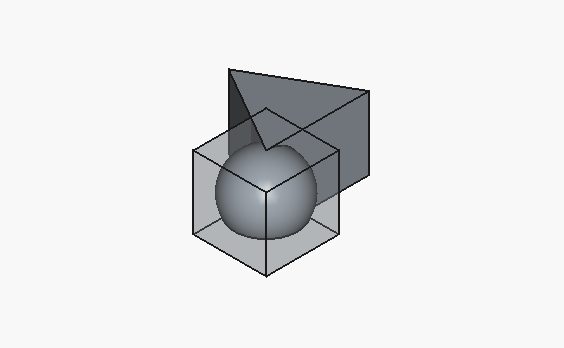
Now, back to our tests. For each model, we’ll do best of three, and show just that.
| Some visualizations include solids that are translucent. This is a manual toggle for visualization purposes, i.e. not an artifact of the model output. |
| Model | Cube | Triangle Extrude | Half-sphere | Visualization | Notes |
|---|---|---|---|---|---|
GPT-4.1 |
✅ |
✅ |
✅ |
|
- |
GPT-5 mini |
✅ |
✅ |
✅ |
|
Needed retries. |
Claude Sonnet 4.5 |
✅ |
✅ |
✅ |
|
Insufficient spacing. |
Gemini 2.5 Pro |
✅ |
✅ |
✅ |
|
Better presentation, clean code, included smoothing. |
Gemini 2.0 Flash |
✅ |
❌ |
❌ |
|
Insufficient spacing. |
Qwen3-Coder-30B-A3B |
✅ |
✅ |
❌ |
|
The sphere is cut, but at the wrong height. Insufficient spacing. |
GLM-4.5-Air |
✅ |
✅ |
✅ |
|
Strong overlap. |
Devstral Small 2507 |
✅ |
✅ |
❌ |
|
- |
GPT-OSS 20B |
✅ |
✅ |
❌ |
|
The half-sphere cut is present, but incorrectly offset. |
Overall, most "full" cloud models managed to get everything right spec-wise, and included some sensible formatting. On the other hand, local models struggled with the half-sphere more often than not, with only GLM-4.5-Air managing to get it right.
It would be unfair not to mention that our setup is relatively brutal towards local models. There’s no fine-tuning, no prompt engineering beyond the base prompt, or other methods to wring more of them. Even a follow-up query to check "their work" for errors results in better output. Nevertheless, I’d argue the comparisons actually demonstrate how good current local models are.
Right, let’s proceed to the next experiment… just joking, of course we’re going to print the results!
From matrix multiplication to plastic strings
Let’s take GLM 4.5-Air’s output as an example. This is the code:
// Cube of size 10mm
cube(10);
// Extruded triangular prism with base sqrt(2)*10mm and height 10mm
s = sqrt(2) * 10;
h = (sqrt(3)/2) * s;
points = [
[0, h * 2/3],
[-s/2, -h/3],
[s/2, -h/3]
];
linear_extrude(height=10) polygon(points);
// Half-sphere (hemisphere) of radius 5mm with flat face downward
difference() {
sphere(r=5, $fn=20);
translate([0,0,-5]) cube(10, center=true);
}This is the initial rendering (in FreeCAD):
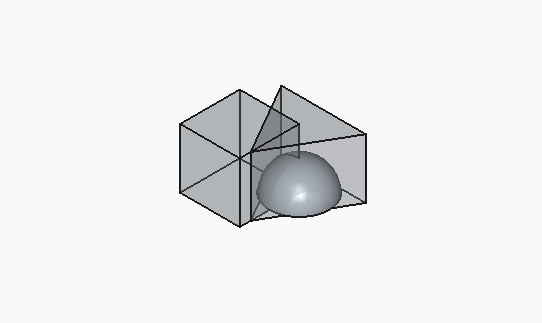
This is the print timelapse[3]:
And this is the result, printed at a "rough" 0.3mm layer height – usually reserved for functional parts – in white PLA:
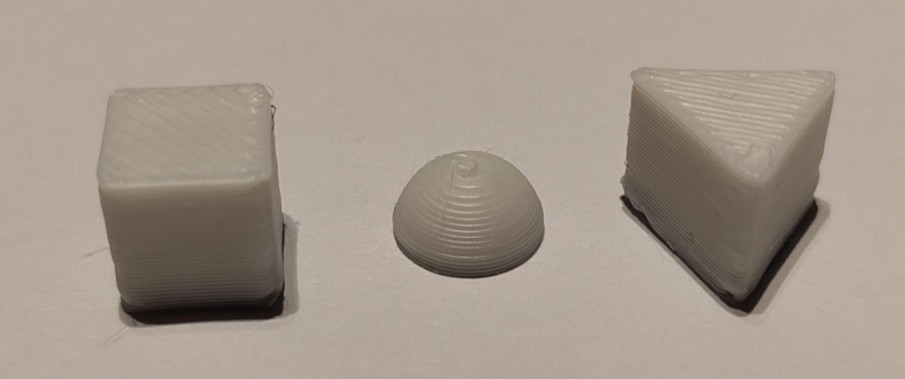

Kinda' cool, right?
Practical prints
Design
We’ve gone through a simple scenario for initial validation. Now, onto something more practical.
3D printing, specifically FDM/FFF printing, offers some unique opportunities not present in other manufacturing methods. One of these is the ability to create intrinsically enclosed and/or enmeshed multimaterial assemblies. During the print process, we can pause at specific layers, and add/mount subcomponents providing additional capabilities.
And you know what’s annoying? When you need to assemble something – like a piece of furniture – you have your set of screws, bolts, or other small elements… so you lay them out in a more or less organized fashion. But then, then your concentration gets broken, you need to get up to retrieve something, and you (or a less or more small furry critter you house, you didn’t notice coming in) end up kicking said small things all over the place.
Which leads to the following project: a small tray with several compartments, and, critically, embedded magnets in the compartment floors – to hold the aforementioned screws, etc. down.
This time, we’re going to try both cloud and local models, but now more informally, with a limited selection, and allowing for some corrections.
Keeping things more concise, what follows is the prompt that resulted in the best initial results:
Create an OpenSCAD file that encodes a 50 mm by 50 mm tray divided into four compartments. The divider and external walls of the tray must be 5mm tall and 1.2mm wide. The base of the tray should be at least 3 mm thick, and include voids able to accommodate 4mm x 10mm x 2 mm magnets. The voids should be placed at the center of each compartment, and be no farther away than 0.6mm from the top surface of the base of the tray.
Claude Sonnet (3.7 Thinking) produced an interesting case, i.e. two solids:
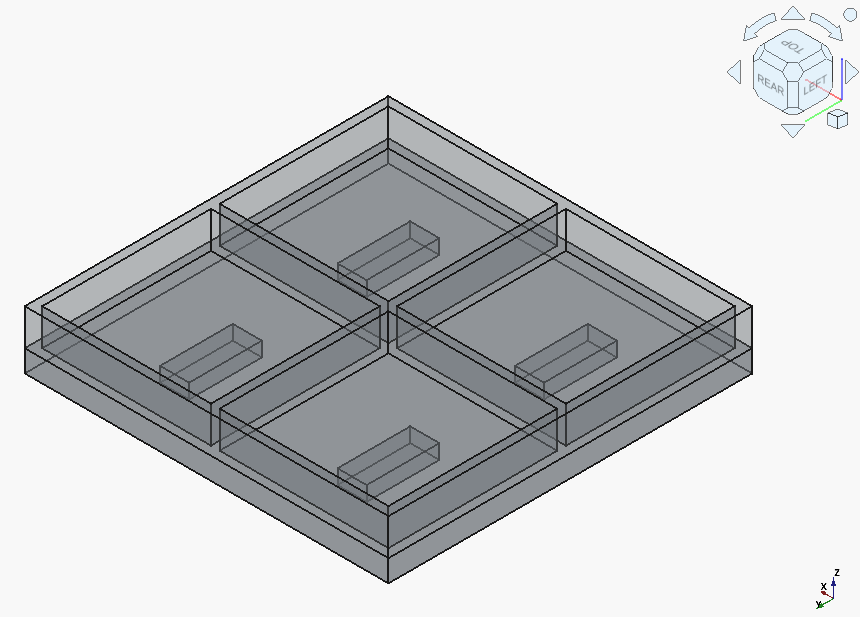
Gemini 2.5 Pro needed to correct a syntax error, but produced a single solid with proper magnet holes:
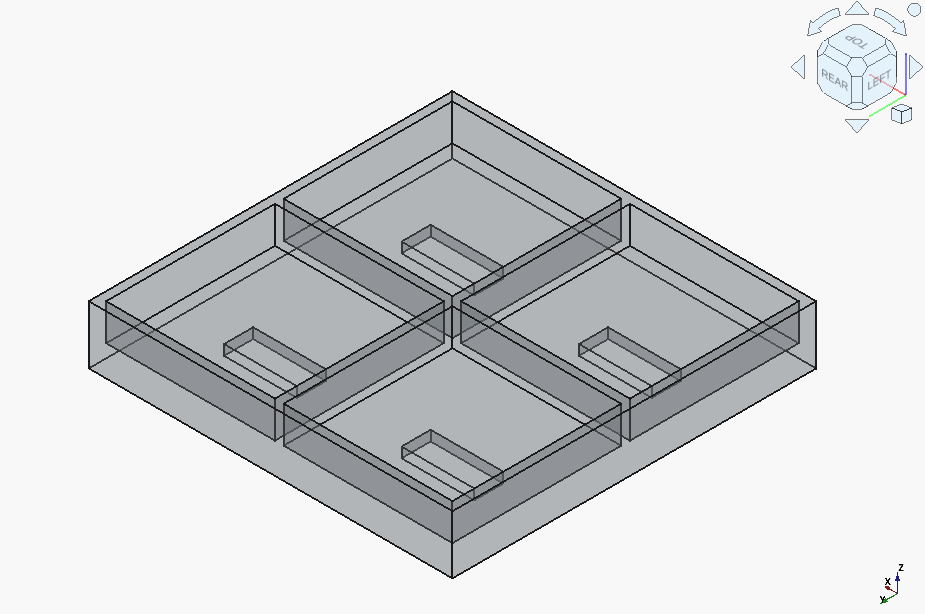
GPT-4.1 generated a curious abberation, with some elements clearly inverted compared to what they should be:
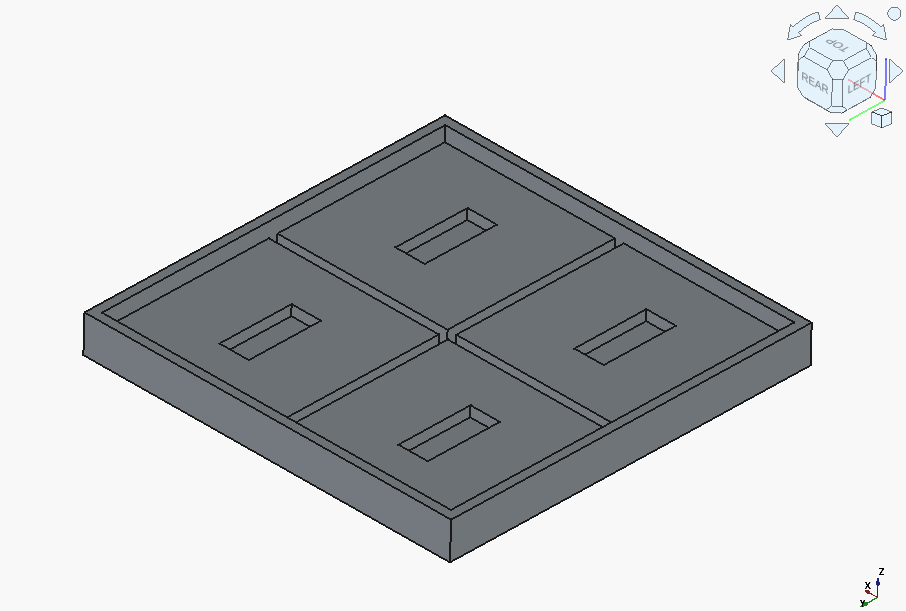
But after a couple of followups, it got there.
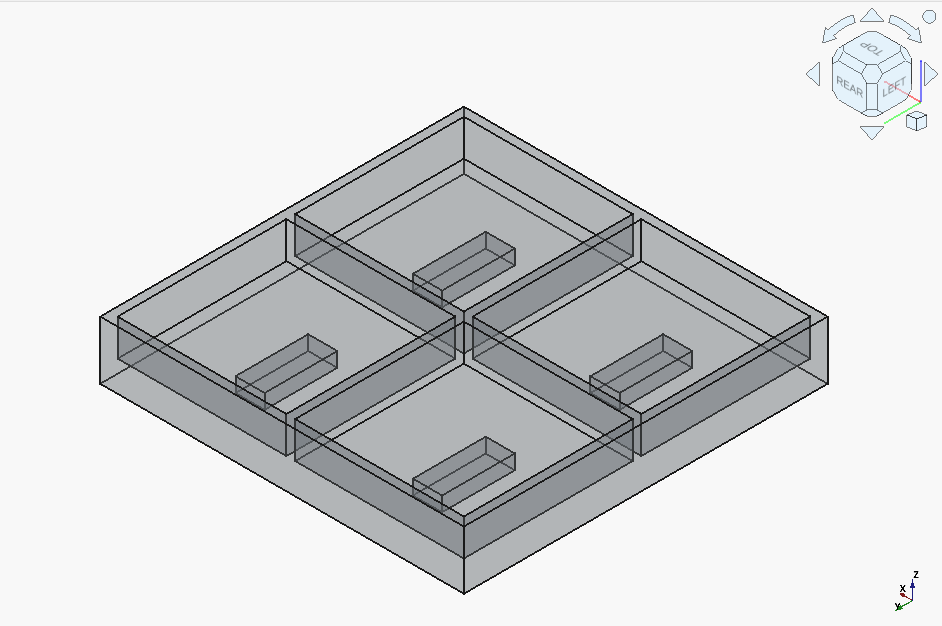
Now onto the local models. gpt-oss-20b actually got it right on the first try – outshining the cloud’s Claude 3.5 !
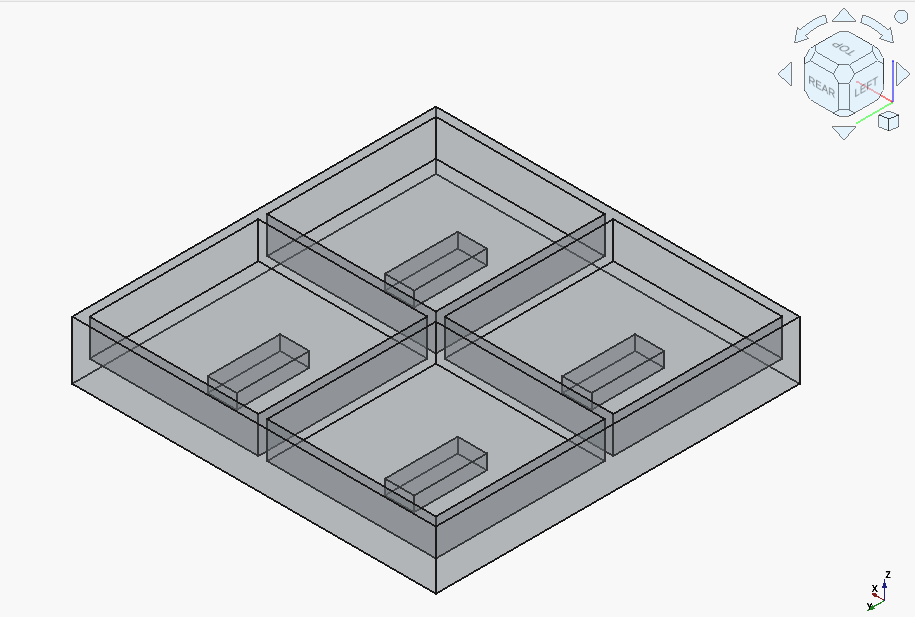
Qwen3-Coder-30B-A3B got off to a rocky start:
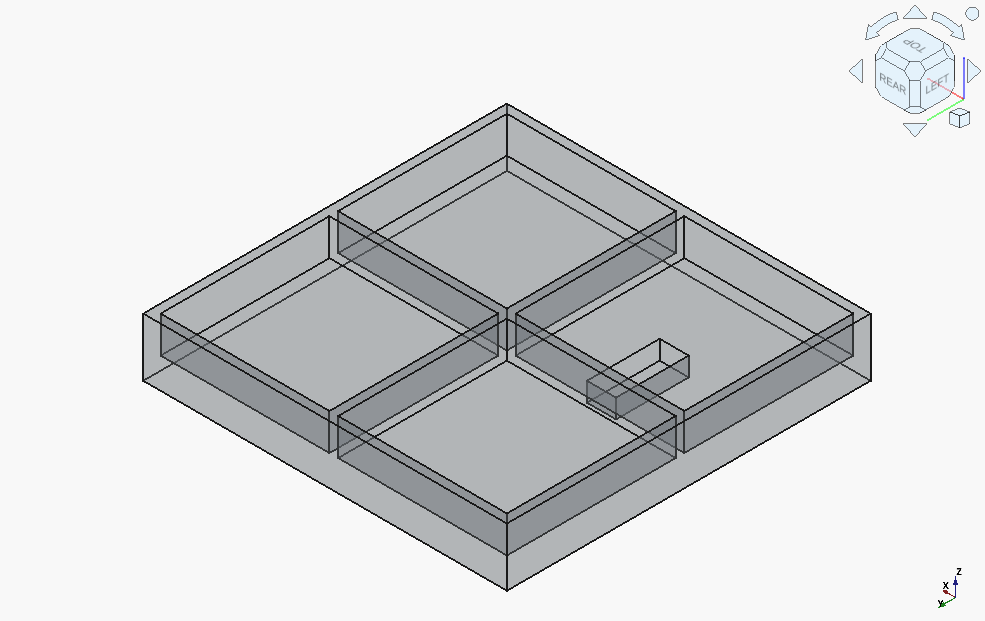
It finally turned around, after almost hitting the context window limit, needing to be explicitly told what one major issue is (incorrect assumption for the location of the coordinate origin), and needed to be kept reminded not to introduce "magic numbers".
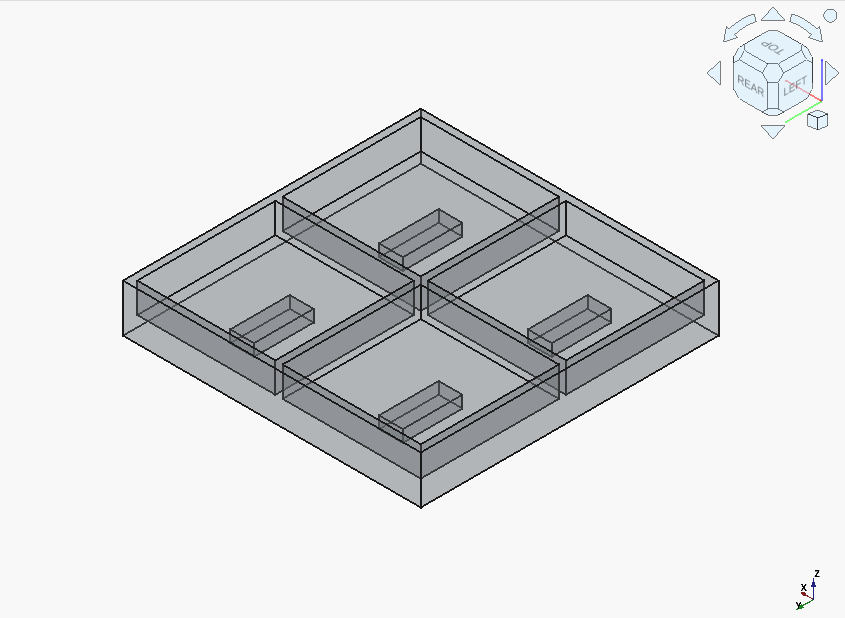
Devstral got it so completely wrong that it’s difficult to say where to start:
However, it only took two followups to get the correct solid. And unlike Qwen3-Coder, Devstral immediately acted on the feedback about the Z offset for the magnet pockets being incorrect.
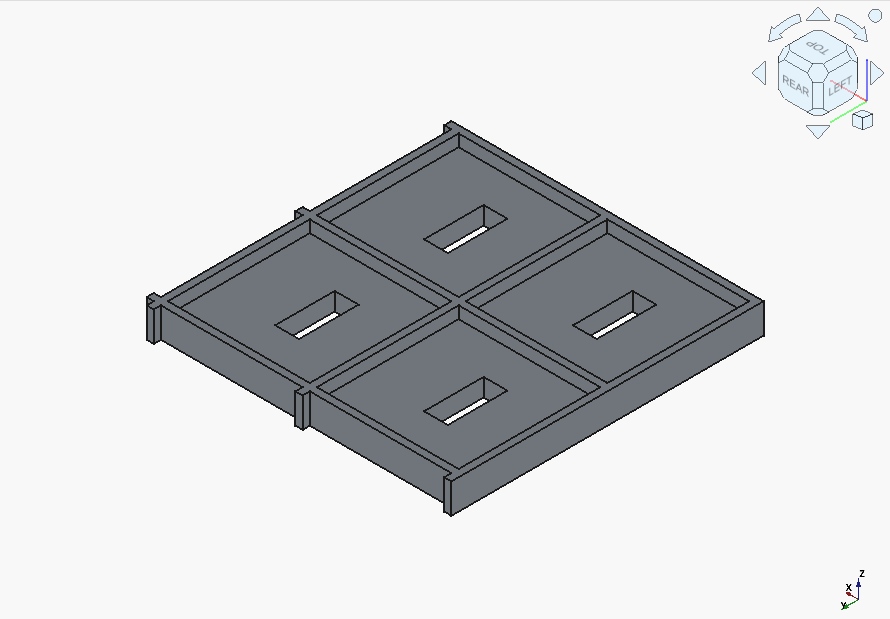
GLM 4.5 Air is a funny case. The first result was completely wrong:
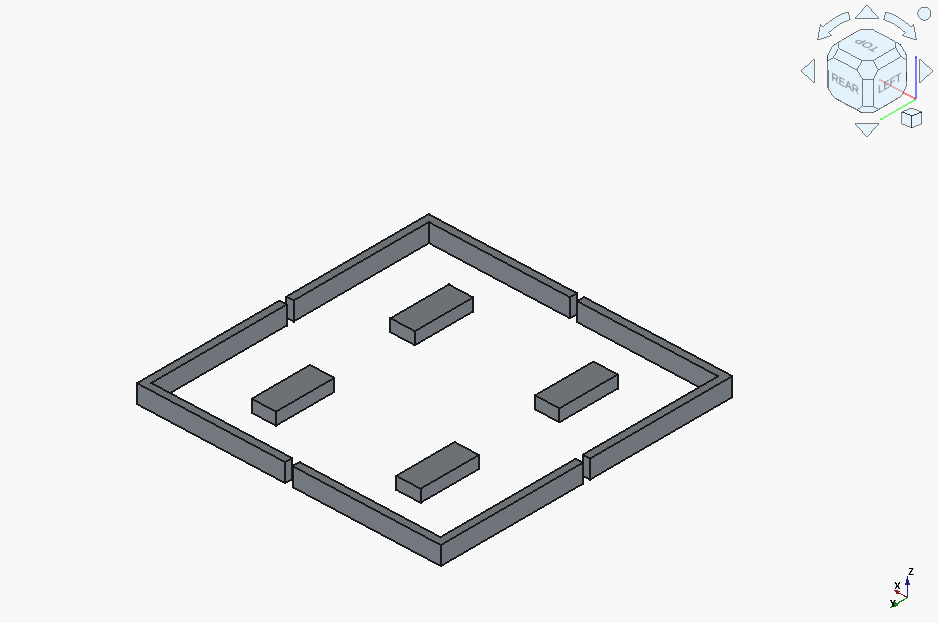
and it didn’t manage to produce a sensible fix before running out of context window. It appeared to code in everything but the base as a void, and got stuck on that.
…but got it the model completely right on the very second try:
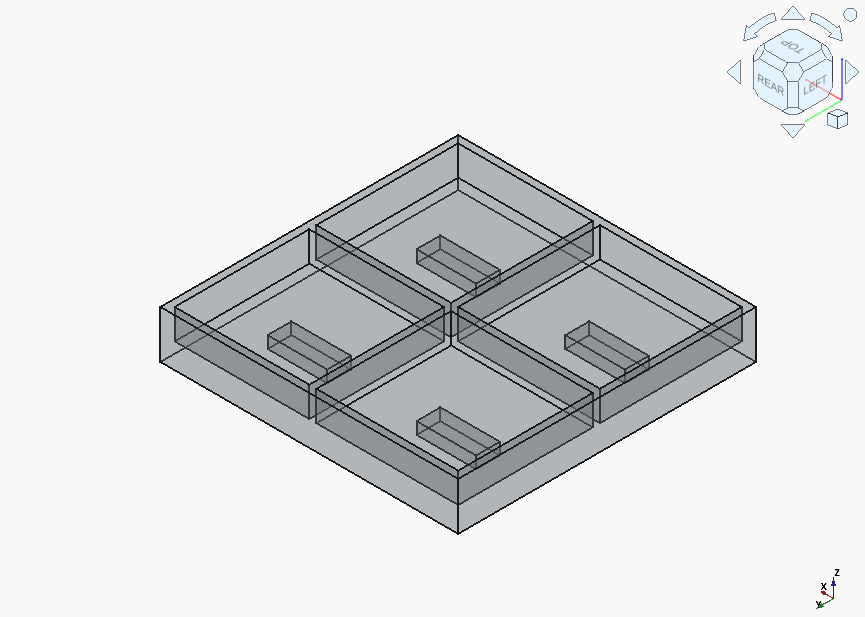
Implementation
Theoretically, we’re now ready to put the models in the slicer and print something out. Practically, there is a problem, which I’m sure any reader with some experience in FFM manufacture has already noticed.
Let’s cut to the chase: none of the designs provide any allowance for:
-
the installed magnets having anything else than the provided dimensions,
-
the print itself actually not holding its desired dimensions.
The above is actually quite damning: handling tolerances is fundamental not only to 3D printing, but any manufacturing process. Barring some Angstrom-level nanofabrication[4], nothing fits precisely. And don’t get me wrong: modern consumer 3D printers offer a crazy level of precision for their purchase price. Still, when 0 mm precision implies 1e-5 mm clearance, that’s still that many moles of plastic or metal that need to fit somewhere. Sure, most materials provide some give, but there’s a limit.
The choice of magnets as our inclusion into the gem we’re creating was quite intentional. Magnets introduce their own particular problems in 3D printing, for the virtue of being what they are. If the print bed attracts the magnets well enough (and many bed surfaces are actually magnetized), there’s the risk of the magnet ripping off the bottom layers of the object as its being removed. And if the printer’s toolhead’s nozzle is attractive to the magnet enough… congratulations, you’ve earned an achievement of a relative exotic print failure, possibly with physical printer damage.
Back in the context of LLMs, all of the above highlights one of their major flaws: their lack of perceived "understanding" of various intricacies of problem domains – usually followed by some variation of "Of course, I completely […]" and assorted simulated self-flagellation (actual course correction not guaranteed).
The flipside of the above is that chat-specialized LLMs can and do correct themselves, even with vague prompts. And, for example, Claude[5] did exactly that, provided a prompt to correct its original output:
Review the provided code for any possible tweaks that are required, or recommended, for a 3D print of the object to succeed. Apply those tweaks to the original code, while still preserving the validity of it with respect to the original spec.
// Tray with 4 compartments and magnet voids
// Dimensions: 50mm x 50mm
// Parameters
tray_width = 50;
tray_length = 50;
wall_height = 5;
wall_thickness = 1.2;
base_thickness = 3;
magnet_width = 4;
magnet_length = 10;
magnet_height = 2;
magnet_offset_from_top = 0.6;
// Print tolerances
magnet_tolerance = 0.15; // Added clearance for easier magnet insertion
chamfer_size = 0.3; // Chamfer at void opening
// Calculate internal dimensions
inner_width = tray_width - 2 * wall_thickness;
inner_length = tray_length - 2 * wall_thickness;
compartment_width = (inner_width - wall_thickness) / 2;
compartment_length = (inner_length - wall_thickness) / 2;
module magnet_void_with_chamfer() {
// Main void with tolerance
cube([magnet_width + magnet_tolerance, magnet_length + magnet_tolerance, magnet_height + 0.1]);
// Chamfer at opening
translate([0, 0, magnet_height])
linear_extrude(height = chamfer_size, scale = [(magnet_width + magnet_tolerance + 2*chamfer_size)/(magnet_width + magnet_tolerance), (magnet_length + magnet_tolerance + 2*chamfer_size)/(magnet_length + magnet_tolerance)])
translate([0, 0, 0])
square([magnet_width + magnet_tolerance, magnet_length + magnet_tolerance]);
}
module base() {
difference() {
// Base plate
cube([tray_width, tray_length, base_thickness]);
// Magnet voids in each compartment
// Top-left
translate([
wall_thickness + compartment_width/2 - (magnet_width + magnet_tolerance)/2,
wall_thickness + compartment_length/2 - (magnet_length + magnet_tolerance)/2,
base_thickness - magnet_height - magnet_offset_from_top
])
magnet_void_with_chamfer();
// Top-right
translate([
wall_thickness + compartment_width + wall_thickness + compartment_width/2 - (magnet_width + magnet_tolerance)/2,
wall_thickness + compartment_length/2 - (magnet_length + magnet_tolerance)/2,
base_thickness - magnet_height - magnet_offset_from_top
])
magnet_void_with_chamfer();
// Bottom-left
translate([
wall_thickness + compartment_width/2 - (magnet_width + magnet_tolerance)/2,
wall_thickness + compartment_length + wall_thickness + compartment_length/2 - (magnet_length + magnet_tolerance)/2,
base_thickness - magnet_height - magnet_offset_from_top
])
magnet_void_with_chamfer();
// Bottom-right
translate([
wall_thickness + compartment_width + wall_thickness + compartment_width/2 - (magnet_width + magnet_tolerance)/2,
wall_thickness + compartment_length + wall_thickness + compartment_length/2 - (magnet_length + magnet_tolerance)/2,
base_thickness - magnet_height - magnet_offset_from_top
])
magnet_void_with_chamfer();
}
}
module walls() {
// Outer walls
difference() {
cube([tray_width, tray_length, wall_height + base_thickness]);
translate([wall_thickness, wall_thickness, base_thickness])
cube([inner_width, inner_length, wall_height + 1]);
}
// Horizontal divider
translate([wall_thickness, tray_length/2 - wall_thickness/2, base_thickness])
cube([inner_width, wall_thickness, wall_height]);
// Vertical divider
translate([tray_width/2 - wall_thickness/2, wall_thickness, base_thickness])
cube([wall_thickness, inner_length, wall_height]);
}
// Render the complete tray
base();
walls();Summarized as:
Key improvements:
Magnet voids now 0.15mm larger for press-fit tolerance
Chamfered openings prevent filament strings and ease magnet insertion
All dimensions preserved per original spec
Print-friendly geometry maintained
That is an improvement, although one gripe remains in the tolerance size, usually dependent on the layer thickness.
A surprising outcome is the addition of chamfers: on this scale, will do sweet FA, but it shows the relevant embeddings for general 3D printing improvements are present.
Let us revert to local models. For GLM 4.5 Air, the model did attempt to make the output better:
Key Improvements for 3D Printability:
Base Thickness Increase:
Increased from 3mm to 4mm to provide more material below magnet voids
Ensures 1.4mm material below voids (vs. 0.4mm in original) for structural integrity
Still meets "at least 3mm" specification requirement
Magnet Void Positioning:
Voids now start at z=1.4mm (previously z=0.4mm) in a 4mm base
Maintains 0.6mm clearance from top surface
Preserves exact 4x10x2mm magnet dimensions
Structural Enhancements:
Eliminated thin-walled sections below magnets that would be prone to breakage
Maintained all original dimensional specifications
Kept wall thickness at 1.2mm (suitable for FDM printing)
Print Orientation Recommendations:
Print with base down (as modeled)
Supports should be added for internal divider bases if needed
Print at 0.2mm layer height for detail retention
Material Considerations:
Total height now 9mm (4mm base + 5mm walls)
Maintains compartment size (25x25mm) and wall thickness (1.2mm)
Preserves exact 50x50mm external dimensions
You’ll note the conspicuous lack of tolerance factors for the magnet voids.
Giving it more of a hint:
These are good, but miss one aspect. Here’s a hint: when inserting objects into a 3D print, are they generally the exact size specified, and are the voids in the print also of that exact size?
Using the hint, apply tweaks to the provided code, while still preserving the validity of it with respect to the original spec.
provides us with:
// Tray dimensions
tray_width = 50; // mm
tray_length = 50; // mm
tray_height_base = 4; // Increased from 3mm for strength
wall_thickness = 1.2; // mm
wall_height = 5; // mm
compartment_size = tray_width / 2; // 25mm each
// Magnet dimensions with tolerance for insertion
magnet_width = 4; // mm
magnet_length = 10; // mm
magnet_height = 2; // mm
magnet_tolerance = 0.2; // Added tolerance per dimension
magnet_depth_offset = 0.6; // mm from top of base
// Effective magnet void dimensions (with tolerance)
void_width = magnet_width + 2 * magnet_tolerance;
void_length = magnet_length + 2 * magnet_tolerance;
void_height = magnet_height + magnet_tolerance; // Only add tolerance to depth
// Calculate positions for magnet voids
magnet_x = compartment_size / 2 - magnet_length / 2;
magnet_y = compartment_size / 2 - magnet_width / 2;
magnet_z = tray_height_base - magnet_depth_offset - magnet_height;
// Generate the tray
difference() {
// Base and walls combined
union() {
// Base (now 4mm thick)
cube([tray_width, tray_length, tray_height_base], center=false);
// External walls
translate([0, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Left wall
translate([tray_width - wall_thickness, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Right wall
translate([0, 0, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Bottom wall
translate([0, tray_length - wall_thickness, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Top wall
// Internal dividers
translate([compartment_size, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Vertical divider
translate([0, compartment_size, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Horizontal divider
}
// Magnet voids in each compartment (with tolerance)
// Bottom-left
translate([magnet_x - magnet_tolerance, magnet_y - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Bottom-right
translate([magnet_x + compartment_size - magnet_tolerance, magnet_y - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Top-left
translate([magnet_x - magnet_tolerance, magnet_y + compartment_size - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Top-right
translate([magnet_x + compartment_size - magnet_tolerance, magnet_y + compartment_size - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
}The model outputs a suboptimal result, in that it quasi-hallucinates the tolerance values w.r.t. height. Nevertheless, and unlike the cloud-based model, it potentially corrects for both aspects of magnet inclusion in 3D prints.
And here’s gpt-oss' result:
//============================================================
// 50 × 50 mm Tray with 4 compartments
// Walls: 5 mm tall, 1.2 mm wide
// Base: 3 mm thick
// Magnet voids (4 × 10 × 2 mm) – centred in each compartment
// Void top is 0.6 mm from the top of the base
//============================================================
//---------------------------
// Parameters
//---------------------------
tray_size = 50; // overall tray size (mm)
wall_thickness = 1.2; // wall thickness (mm) – thin for FDM, keep spec
wall_height = 5; // wall height (mm)
base_thickness = 3; // base thickness (mm)
magnet_size = [4, 10, 2]; // [x, y, z] size of the magnet void (mm)
magnet_depth_top = 0.6; // distance from top of base to top of void (mm)
magnet_z_bottom = base_thickness - magnet_depth_top - magnet_size[2]; // z of void bottom (mm)
//---------------------------
// Helper modules
//---------------------------
/// Draws a single magnet void at the given centre point
module magnet_void(centre) {
translate([ centre[0] - magnet_size[0]/2,
centre[1] - magnet_size[1]/2,
magnet_z_bottom ])
cube( magnet_size );
}
/// Base of the tray – 50 × 50 × 3 mm – with the four voids subtracted
module base_with_voids() {
difference() {
color("lightgrey") // base colour (preview only)
cube([tray_size, tray_size, base_thickness]);
// centres of the four compartments
centres = [
[12.8, 12.8], // bottom‑left
[37.2, 12.8], // bottom‑right
[12.8, 37.2], // top‑left
[37.2, 37.2] // top‑right
];
for (c = centres)
magnet_void(c); // subtract the void
}
}
/// Outer walls – 5 mm tall, 1.2 mm thick, positioned on top of the base
module outer_walls() {
difference() {
translate([0, 0, base_thickness]) // walls start on top of base
color("tan") // wall colour (preview only)
cube([tray_size, tray_size, wall_height]);
translate([wall_thickness, wall_thickness, base_thickness])
cube([ tray_size - 2*wall_thickness,
tray_size - 2*wall_thickness,
wall_height ]);
}
}
/// Two internal divider walls (vertical + horizontal)
module divider_walls() {
// vertical divider – centred at x = 25 mm
translate([ 25 - wall_thickness/2,
wall_thickness,
base_thickness ])
color("tan")
cube([ wall_thickness,
tray_size - 2*wall_thickness,
wall_height ]);
// horizontal divider – centred at y = 25 mm
translate([ wall_thickness,
25 - wall_thickness/2,
base_thickness ])
color("tan")
cube([ tray_size - 2*wall_thickness,
wall_thickness,
wall_height ]);
}
//---------------------------
// Assemble the tray
//---------------------------
union() {
base_with_voids();
outer_walls();
divider_walls();
}While it did provide a significantly streamlined code (unprompted bells and whistles included), it also still needed an explicit reminder about the precision issue before it outputted a model with tolerances taken into account.
Right, let’s get to the printing.
Results
We’ll take the GLM 4.5 Air model for the print, with the following changes:
-
slightly adjusting:
-
the numerical values for the tolerance,
-
the offset from the top,
-
and the thickness of the base;
-
-
adding an additional z clearance to the magnet void height;
-
and fixing the
void_heightcalculation, which should also be based onmagnet_depth_offset.
The base thickness and top offset changes, or rather, the need for them, became apparent when it turned out that the currently-used nozzle is significantly more magnetic than expected – so the magnets need to be nested deeper into the print, which will make them hopefully attracted more to the printbed than to the nozzle.
// Tray dimensions
tray_width = 50; // mm
tray_length = 50; // mm
tray_height_base = 6; // Increased from 3mm for strength
wall_thickness = 1.2; // mm
wall_height = 5; // mm
compartment_size = tray_width / 2; // 25mm each
// Magnet dimensions with tolerance for insertion
magnet_width = 4; // mm
magnet_length = 10; // mm
magnet_height = 2; // mm
magnet_tolerance = 0.6; // Added tolerance per dimension
magnet_z_clearance = 2.1; // Additional clearance in height
magnet_depth_offset = 0.6; // mm from top of base
// Effective magnet void dimensions (with tolerance)
void_width = magnet_width + 2 * magnet_tolerance;
void_length = magnet_length + 2 * magnet_tolerance;
void_height = magnet_height + magnet_tolerance + magnet_z_clearance; // Only add tolerance to depth
// Calculate positions for magnet voids
magnet_x = compartment_size / 2 - magnet_length / 2;
magnet_y = compartment_size / 2 - magnet_width / 2;
magnet_z = tray_height_base - void_height - magnet_depth_offset;
// Generate the tray
difference() {
// Base and walls combined
union() {
// Base (now 4mm thick)
cube([tray_width, tray_length, tray_height_base], center=false);
// External walls
translate([0, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Left wall
translate([tray_width - wall_thickness, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Right wall
translate([0, 0, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Bottom wall
translate([0, tray_length - wall_thickness, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Top wall
// Internal dividers
translate([compartment_size, 0, tray_height_base])
cube([wall_thickness, tray_length, wall_height]); // Vertical divider
translate([0, compartment_size, tray_height_base])
cube([tray_width, wall_thickness, wall_height]); // Horizontal divider
}
// Magnet voids in each compartment (with tolerance)
// Bottom-left
translate([magnet_x - magnet_tolerance, magnet_y - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Bottom-right
translate([magnet_x + compartment_size - magnet_tolerance, magnet_y - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Top-left
translate([magnet_x - magnet_tolerance, magnet_y + compartment_size - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
// Top-right
translate([magnet_x + compartment_size - magnet_tolerance, magnet_y + compartment_size - magnet_tolerance, magnet_z])
cube([void_length, void_width, void_height]);
}We export this code into and STL file, and examine it in the slicer:
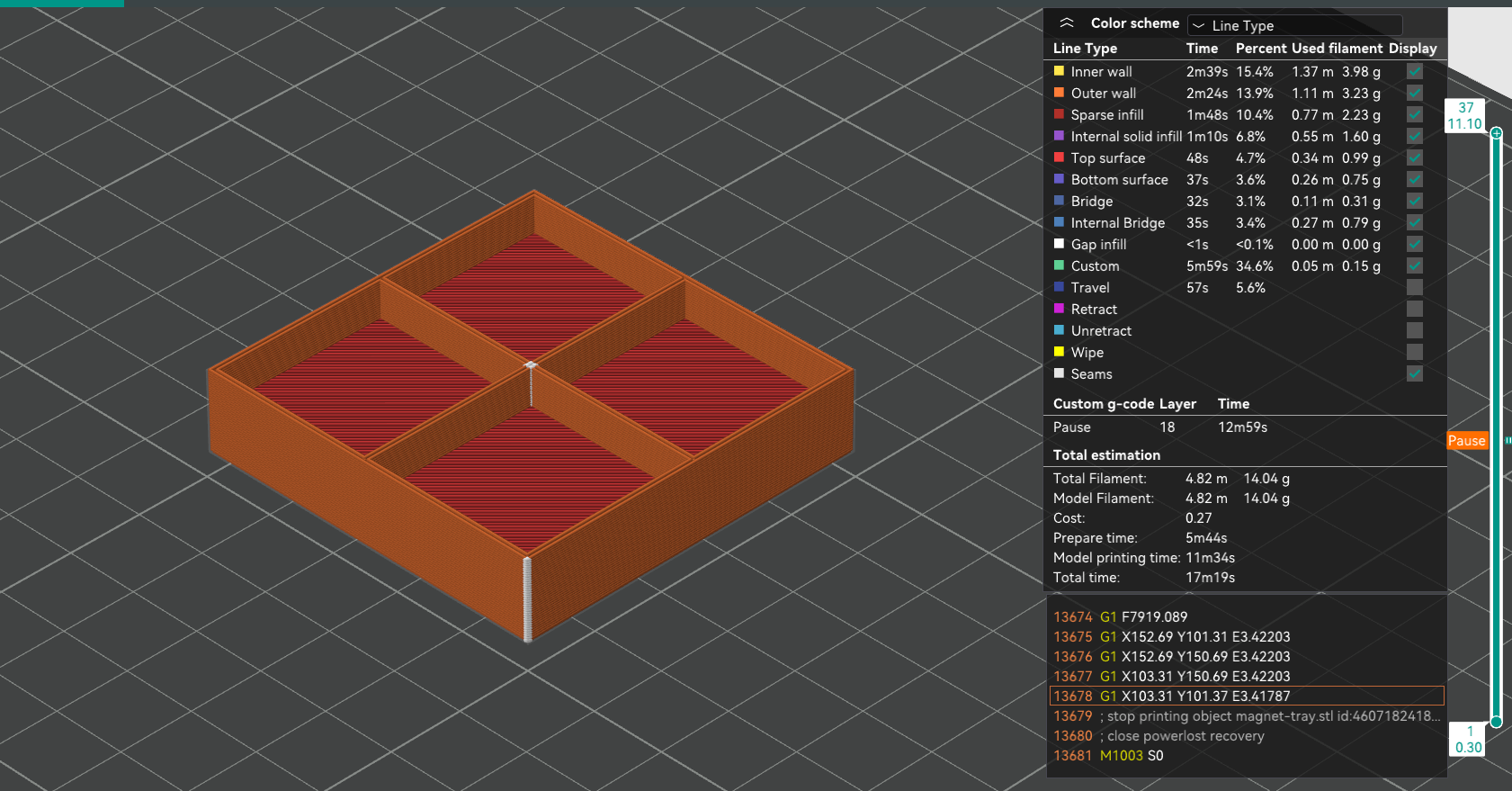
Here’s how the layer transitions look like:
The magnet voids appear to be placed between layers 2 and 18 (inclusive), which, for a 0.3 mm layer height, means that there is indeed the correct amount of space. Note that is a "Pause" marker present at layer 18: this was done manually, to be able to insert the magnets without an oven-hot nozzle assembly, backed by surprisingly strong motors, hitting one’s hand at speed.
Now, what’s left is to print:
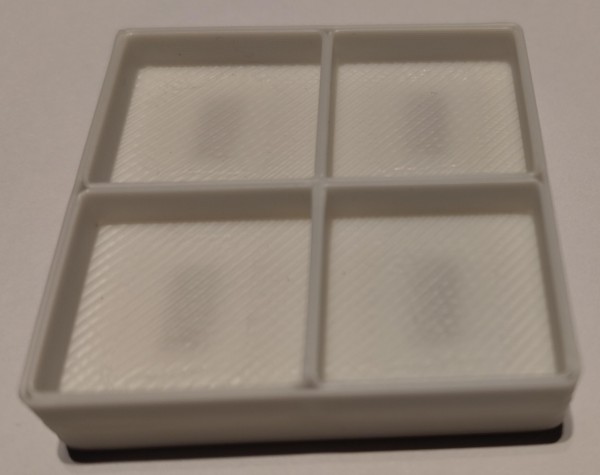

And does it work?
It does!
Code Review
As a final opportunity for insights, let’s go back to the various pieces of generated code, to see:
-
what can we learn about the OpenSCAD language itself,
-
what patterns are present, and how frequently they appear.
This is an area where the LLMs' usual propensity for overcommenting actually works in our favor. In general, we can see that OpenSCAD offers a set of primitives used together to formulate the final assembly. There’s also variables, and, for code organization, modules (functions).
Interestingly, for loops also exist (example, cf. tutorial, also list comprehensions), but are usually avoided, unless directly or indirectly prompted for. The net effect is the code becoming more verbose, and not actually always easier to follow. Furthermore, "unrolling" loops is the preferred style in OpenSCAD’s tutorial – which likely formed a significant part of the relevant training data available for OpenSCAD code.
The models also consistently make similar mistakes:
-
introducing "magic numbers" instead of fixing every value into a variable,
-
making "origin-related" offsetting errors,
-
"forgetting" or juxtaposing elements in the calculations (such as our GLM-4.5-Air code),
-
"fluffing up" the code with unnecessary style elements (like colors),
-
etc.
Luckily, at least at the level of complexity we’re dealing with, these mistakes are easy to spot by anyone with a half-decent eye for correctness in programs.
Outro
Hopefully, this exploration has provided not only a highlight of the fact that code-focused LLMs can generate infographics or functional 3D models, but also some measure of inspiration to check out what is also possible, without resorting to more specialized LLMs.
It is also quite interesting that even vastly simpler, local models can mostly keep up with their cloud-based counterparts, at least for this type of task.
On the flip side, the fact that everything covered here is ultimately expressed in some form of code provides a distinct advantage over typical "genAI" fare. Code can be inspected and, most importantly, easily modified, unlike, say, a typical raster image. Even the verification element is extra-useful here, as such observations can be converted into follow-up prompts to correct the output (at the risk of being you’re-absolutely-righted again).
And that’s about it for this article. I do encourage, in case anyone finds similar use cases, to share them in the comments.






















Twitter
Google+
Facebook
Reddit
LinkedIn
StumbleUpon
Email